Generative AI and NLME
2025-10-12
The Connection Between NLME and Generative AI
Generative AI
Goal: Generate data indistinguishable in distribution to real data
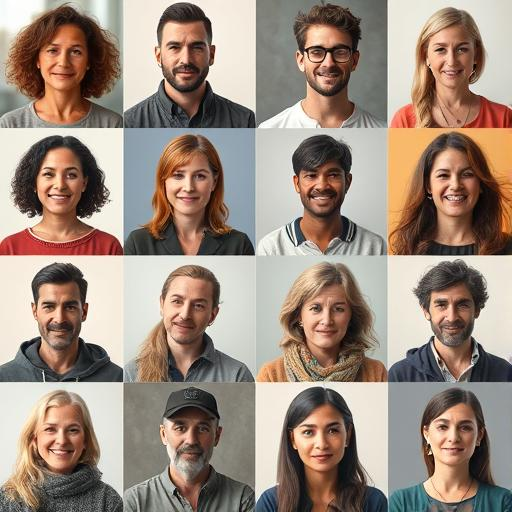 ~
~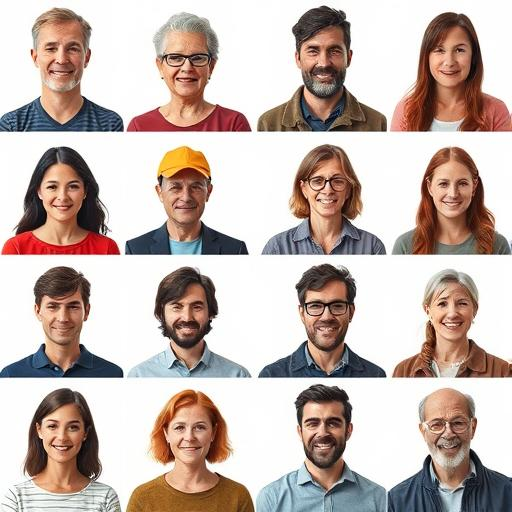 
www.thispersondoesnotexist.com
How is that done?
How is that done?
Generative models
- Definitions
- \(z\): latent variables of dimension \(d\)
- \(y\): observed data
- \(y_g\): generated/simulated/synthetic data
- Model
\[ \begin{aligned} y_g &= f(z) + \epsilon \\ z &\sim Normal(0, I_{d\times d}) \\ \epsilon &\sim Normal(0, \sigma^2) \end{aligned} \]
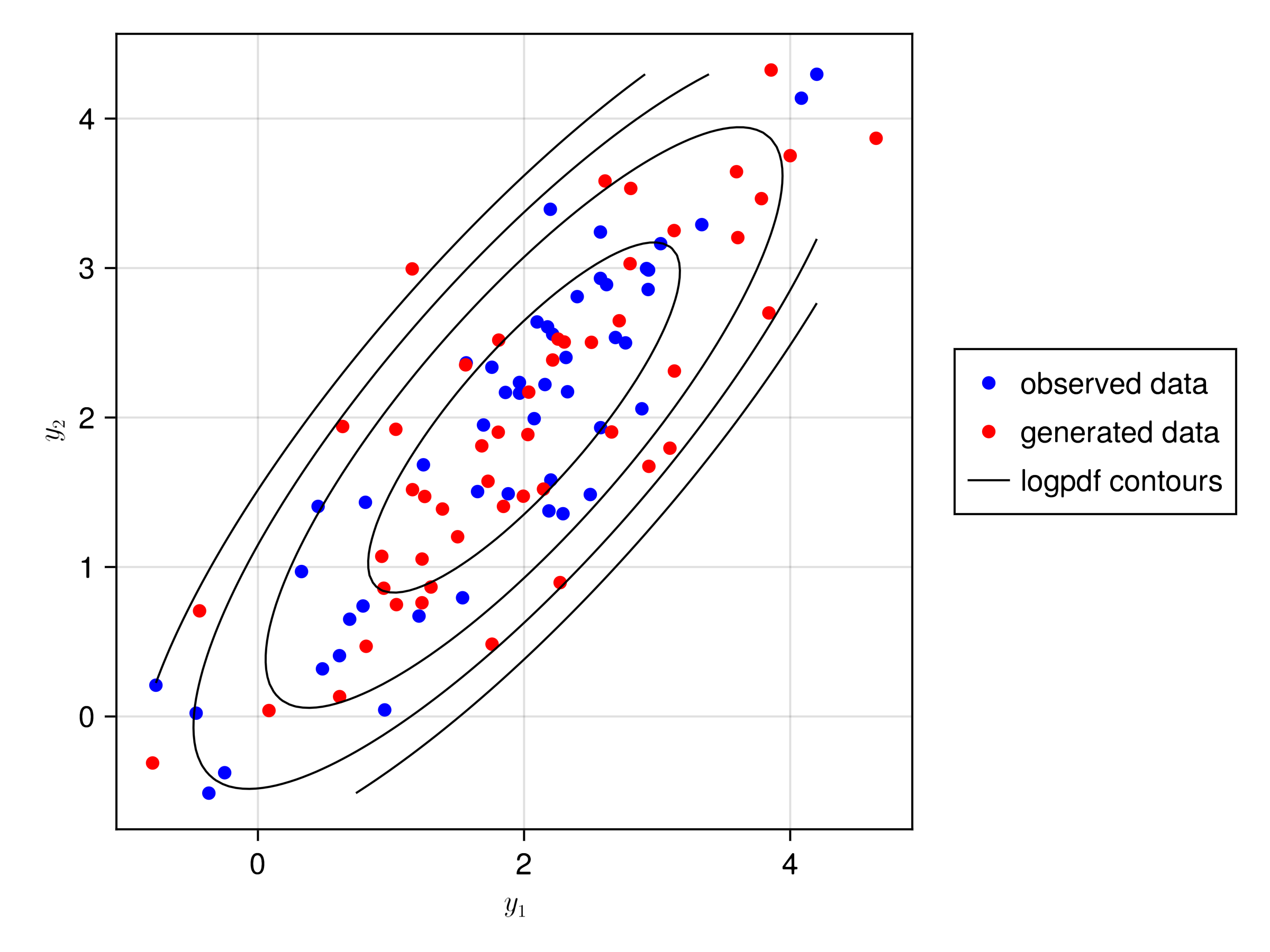
- Objective: find \(f\) such that the distribution of \(y_g\) is close to the distribution of the observed data \(y\)
NLME is Generative AI!
- Definitions
- \(\eta\): latent variables of dimension \(d\) and covariance matrix \(\Omega\)
- \(x\): observed covariates
- \(dv\): observed data/response
- \(dv_g\): generated/simulated/synthetic data
- Model
\[ \begin{aligned} dv_g &= f_\theta(\eta, x) + \epsilon \\ \eta &\sim Normal(0, \Omega) \\ \epsilon &\sim Normal(0, \sigma^2) \end{aligned} \]
- Objective: find \(f\) such that the conditional distribution of \(dv_g | x\) is close to the distribution of the observed data \(dv\)
NLME is Generative AI!
NLME objective: Maximize marginal likelihood of observations \(y\) given covariates \(c\):
\[ p_\theta(y | c) = \int p_\theta(y | \eta, c) \cdot p(\eta) d\eta \]
Variational Autoencoder (GenAI) objective: Maximize likelihood of data \(x\):
\[ p_\theta(x) = \int p_\theta(x | z) \cdot p(z) dz \]
They’re identical!
- Random effects \(\eta\) ↔︎ Latent variables \(z\)
- Observations \(y\) ↔︎ Generated data \(x\)
- Individual predictions ↔︎ Generative model
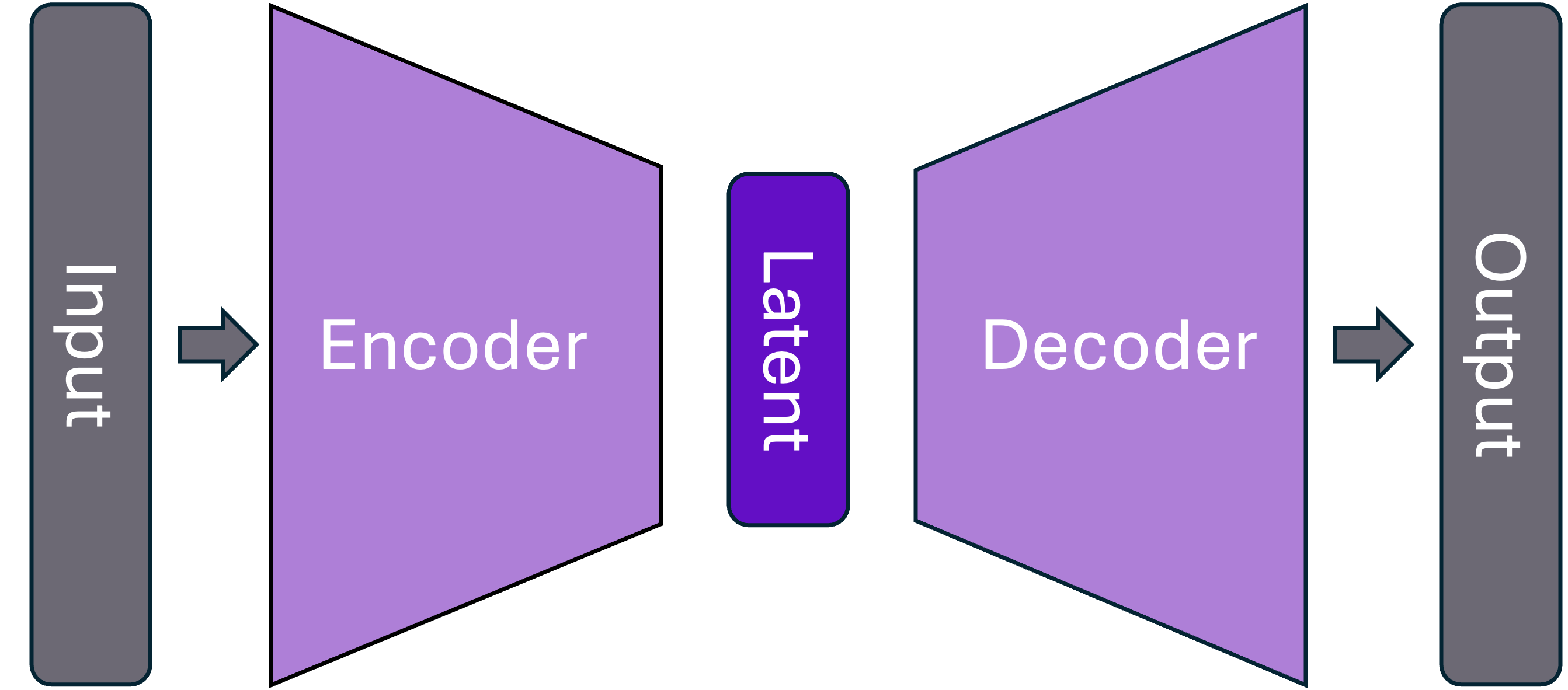
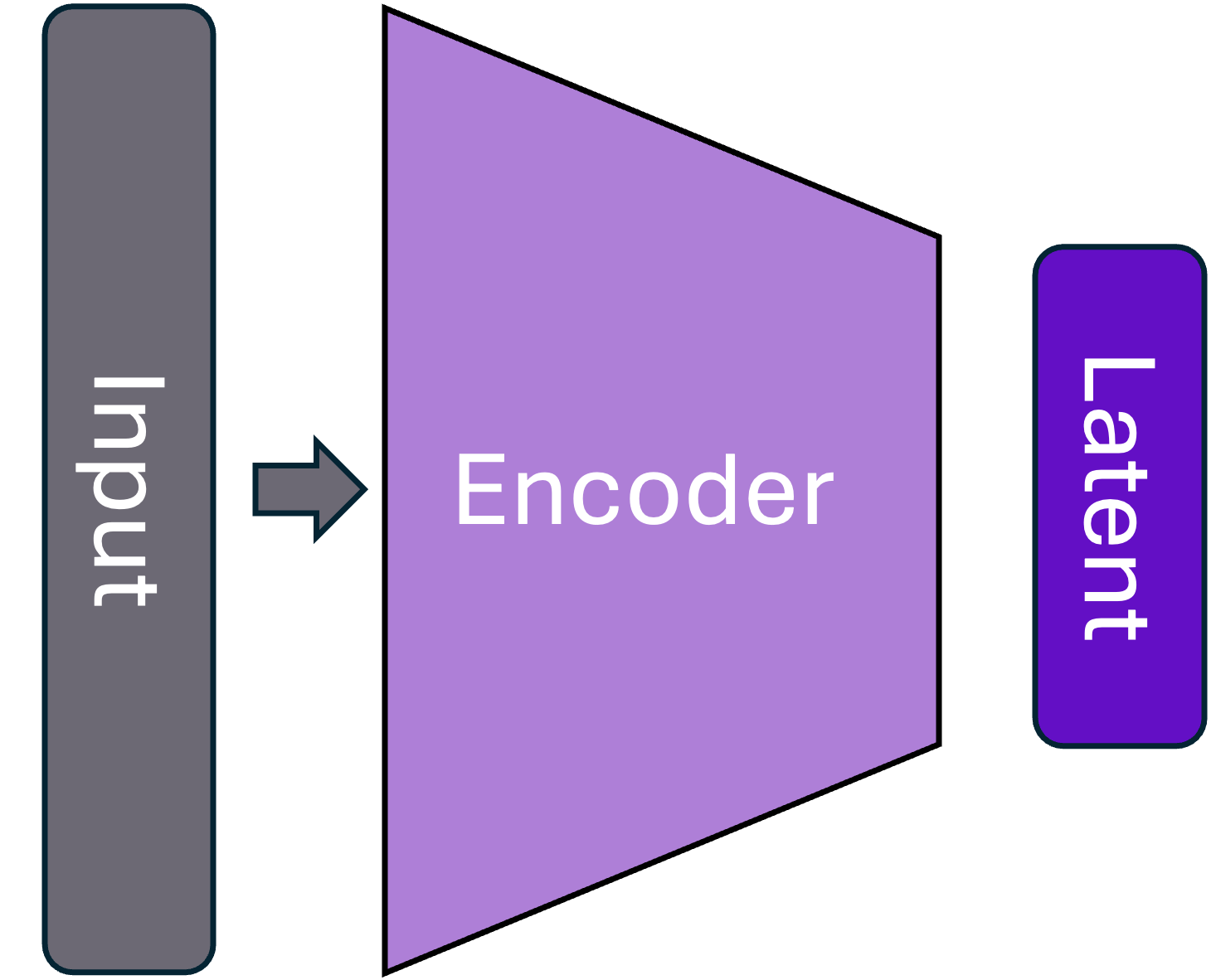
Generative AI – Typical anatomy
NLME as GenAI
What Are Latent Variables?
In Traditional NLME: Meaning is structurally engineered
\[ CL = tvCL \cdot e^{\eta_1} \]
In DeepNLME: More flexible, but still some structure
\[ \frac{dR}{dt} = NN\left(\frac{Central}{Vc}, R, \eta_1, \eta_2 \right) \]
In Pure GenAI: Meaning emerges from data and training
\[ p_\theta(x) = \int p_\theta(x | z) \cdot p(z) dz \]
\(z\) captures informative properties not directly observed
Latent Variables as Information
For images:
- Not pixel-by-pixel intensity
- Rather: Objects, characteristics, actions, style, lighting
For text:
- Not individual words
- Rather: Sentiment, information content, writing style, language
For clinical data:
- Not individual measurements
- Rather: Disease state, treatment response, patient phenotype