Epidemiology Demo
DeepNLME for Population-Level Viral Dynamics
Niklas Korsbo
2025-10-12
Setup
Epidemiological Modeling Challenge
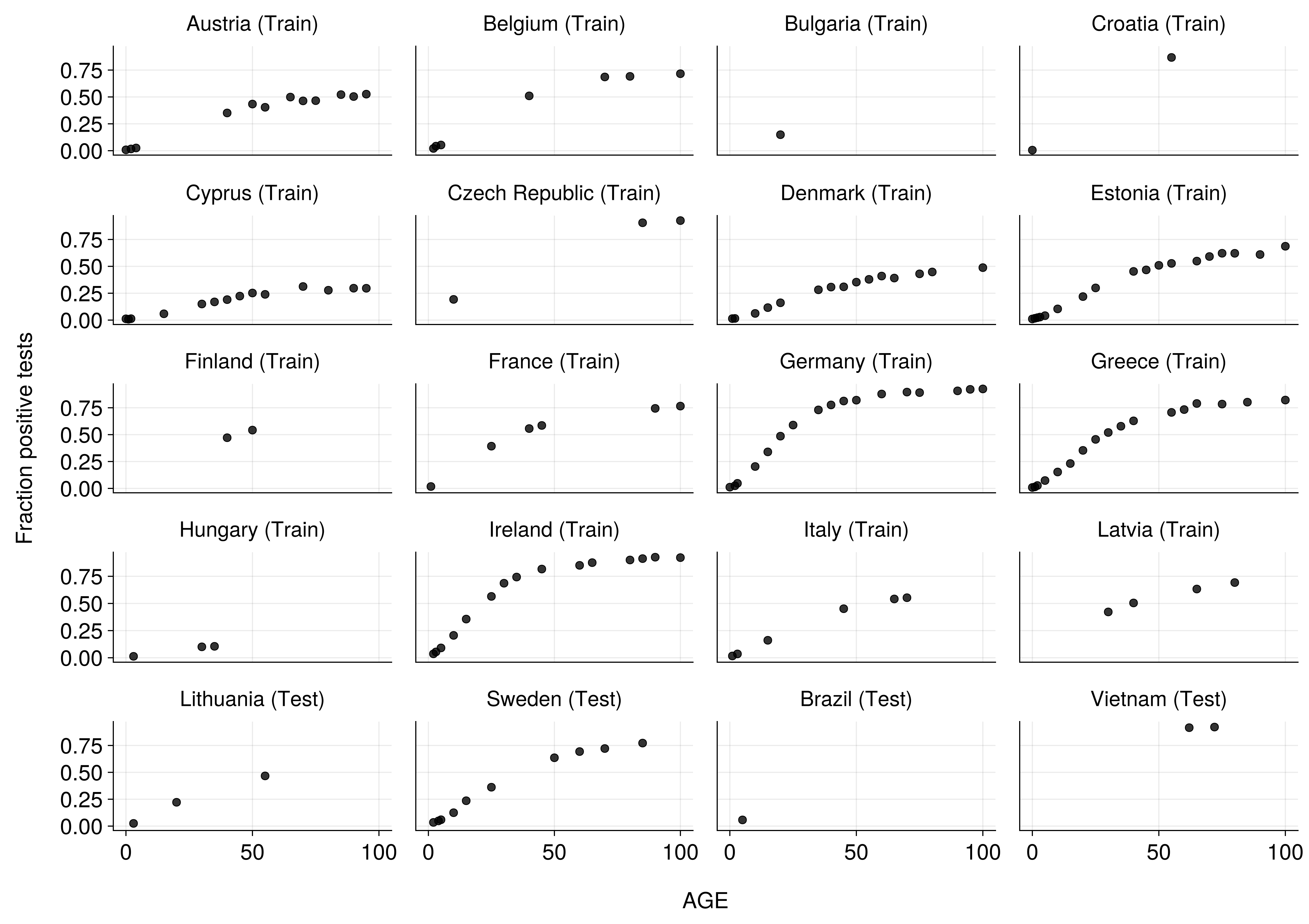
The Problem: Age-Stratified Seroprevalence
The Challenge
- Multi-country seroprevalence data across age groups
- Unknown force of infection λ(age) patterns
- Heterogeneous data quality: Some countries well-sampled, others sparse
- Need to share information between heterogeneous countries
- Discover age-specific transmission patterns
DeepNLME approach: Let data discover the functional form!
DeepNLME Epidemiological Model
SIR model, except that we don’t need the I or R here.
Key features:
- Neural network λ discovers age-specific force of infection
- Individual random effects η allow country-specific scaling
- Shared function across populations with individual tuning
- Mechanistic structure preserves epidemiological interpretation
Results
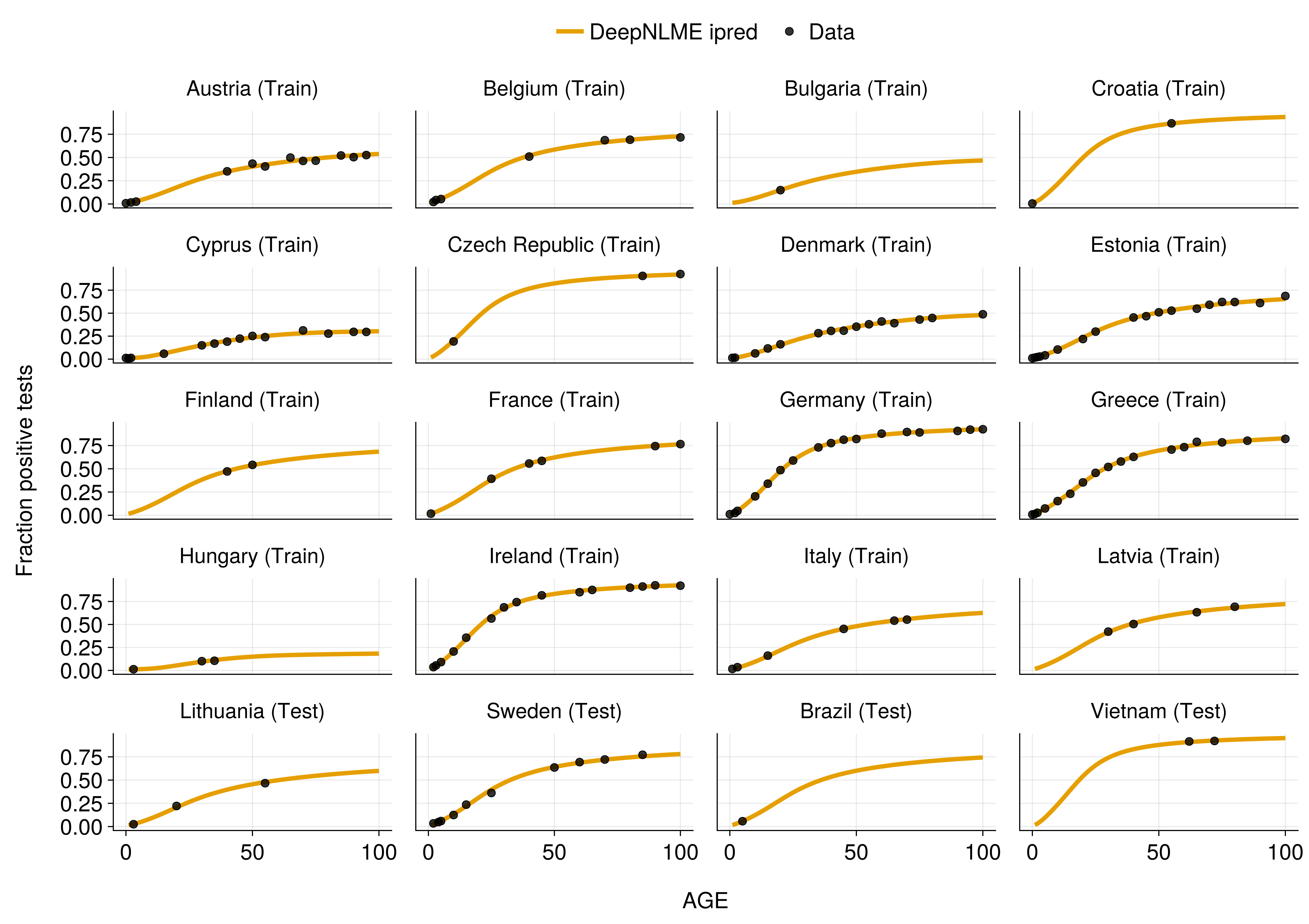
Model Predictions
Excellent fit across all countries, including sparsely sampled test data!
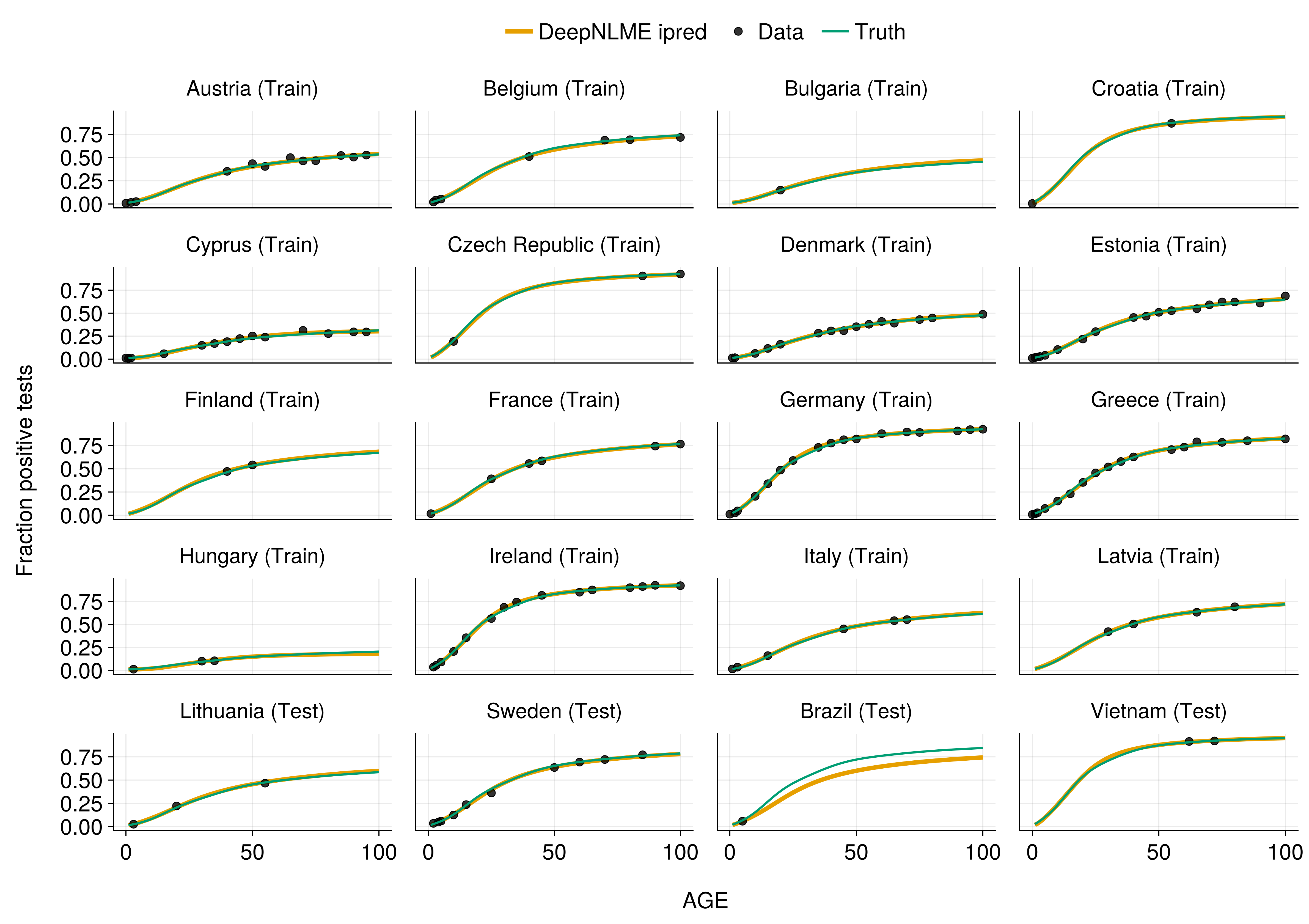
Comparison with Truth
Neural network accurately recovers the true underlying force of infection!
Discovered Force of Infection Pattern
Complex multi-modal pattern discovered automatically from data!
Information Sharing Across Populations
Dense data informs sparse data through shared neural network function!
Key Insights
What DeepNLME Achieved
- Data-driven discovery of force of infection λ(age)
- Automatic pattern recognition without pre-specifying functional forms
- Information sharing
- Individual adaptation / posterior predictions
- Robust extrapolation to poorly sampled regions/countries